I'm just going to show some recent work on extending the Polarizable Embedding from being a focused QM/MM model into an approach to generate parameters for entire proteins or DNA. Here is a snapshot of the human protein GB3 (PDB: 2OED) with the central QM-fragment we are interested in - here LYS4 - shown in vdw-spheres. Also in the QM-region are neighbouring residues and stuff (side-chains and water) that binds to the central fragment of interest) all highlighted in cyan in sticks. The rest of the protein along with a lot of water is the MM-region. Only the protein is shown in a classical cartoon rendering.
Enjoy
Showing posts with label polarizable embedding. Show all posts
Showing posts with label polarizable embedding. Show all posts
Friday, November 8, 2013
Friday, August 23, 2013
Be careful when you extrapolate to the complete basis set limit!
In our soon-to-be-finised manuscript on shielding constants obtained through the QM/MM with the MM-region described by a high-quality polarizable embedding potential (PE), I've done a thing I should have done a while ago. But, as it is always the case with these things, you end up doing it in the wrong order.
How do you continue from these values to an estimate of the basis set limit at infinity? According to Jans blog post, you should fit your data according to
$$
Y(l_{max}) = Y_{CBS} + a \cdot e^{b \cdot l_{max}}
$$
to get the most accurate result. I have done so in using WolframAlpha and I have obtained the following plot (blue dashed line) with $Y_{CBS}=-176.5$ ppm.
"That is a terrible fit!", I can hear you say. And indeed it is. What is going on? It turns out (and please give me some references if you know them!) that the pcS-0 results are actually too bad to be taken seriously. The single zeta basis set is not enough to even get a qualitatively correct description of that wave function, i.e. what you get is just wrong. If you remove that point you get the solid blue curve which is a really good fit with $Y_{CBS}=-187.7$ ppm.
If you want to use the alternative extrapolation scheme that Jan provides, i.e.
$$
Y(l_{max}) = Y_{CBS} + b\cdot l_{max}^{-3}
$$
one obtains the red solid curve with $Y{CBS}=-182.9$ ppm which 5 ppm off from the exponential fit.
As Frank (and Grant) are commenting below, one should not trust numbers from SZ basis sets and +Anders Steen Christensen noted that even the DZ results could/should be disregarded. The only problem is that the size of my calculations are increasing, and a 5Z calculation is pretty much out of reach beyond the TZ basis set. Jans post does mention that one should use a DZ quality basis set, shame on me I guess for even trying with pcS-0.
Just be careful out there and remember to extrapolate!
edit1: fixed the last formula
edit2: Frank Jensen has given a lengthy comment on the matter which gives a lot of insigth. Read it below. I've added some clarifying text here and there based on his comments and will likely follow up on it in a later blog post.
I read the blogpost on basis set extrapolation by +Jan Jensen and thought I should try. Here is what I got by calculating the shielding constants for oxygen-17 in acrolein using KT3/pcS-n//B3LYP/aug-cc-pVDZ solvated by a 12 Å sphere of explicit water molecules described by the PE potential.
| pcS-0 | pcS-1 | pcS-2 | pcS-3 |
|---|---|---|---|
| -309.2 | -216.9 | -191.6 | -188.2 |
How do you continue from these values to an estimate of the basis set limit at infinity? According to Jans blog post, you should fit your data according to
$$
Y(l_{max}) = Y_{CBS} + a \cdot e^{b \cdot l_{max}}
$$
to get the most accurate result. I have done so in using WolframAlpha and I have obtained the following plot (blue dashed line) with $Y_{CBS}=-176.5$ ppm.
"That is a terrible fit!", I can hear you say. And indeed it is. What is going on? It turns out (and please give me some references if you know them!) that the pcS-0 results are actually too bad to be taken seriously. The single zeta basis set is not enough to even get a qualitatively correct description of that wave function, i.e. what you get is just wrong. If you remove that point you get the solid blue curve which is a really good fit with $Y_{CBS}=-187.7$ ppm.
If you want to use the alternative extrapolation scheme that Jan provides, i.e.
$$
Y(l_{max}) = Y_{CBS} + b\cdot l_{max}^{-3}
$$
one obtains the red solid curve with $Y{CBS}=-182.9$ ppm which 5 ppm off from the exponential fit.
As Frank (and Grant) are commenting below, one should not trust numbers from SZ basis sets and +Anders Steen Christensen noted that even the DZ results could/should be disregarded. The only problem is that the size of my calculations are increasing, and a 5Z calculation is pretty much out of reach beyond the TZ basis set. Jans post does mention that one should use a DZ quality basis set, shame on me I guess for even trying with pcS-0.
Just be careful out there and remember to extrapolate!
edit1: fixed the last formula
edit2: Frank Jensen has given a lengthy comment on the matter which gives a lot of insigth. Read it below. I've added some clarifying text here and there based on his comments and will likely follow up on it in a later blog post.
Etiketter:
extrapolation,
NMR,
polarizable embedding,
wolfram alpha
Sunday, July 21, 2013
Oxygen, you little rascal! Part 2.
Background
In the ongoing saga (read part I) with acrolein solvated in water, we were having problems with our high quality embedding potential just not converging nicely when increasing system sizes, QM-region sizes. We were using a functional to get good shielding constants (the KT3 functional) with a basis set designed to get good shielding constants (the apcS-1 basis set) but no matter what tried, the widely adopted TIP3P potential for water just did it better than we did. And we were just confused!
Solution
The problem turned out to be the basis set that with the diffuse functions leaked electron density out into the MM-region when using the PE potential and this was causing some disturbance with the induced dipoles. On the contrary, our belief is that TIP3P is a much too soft potential and there is a large degree of error cancellation because of that. Both are effects I would think to be very small and have no real impact but I had also forgotten that we were talking about nuclear shielding constants which are sensitive as f*ck the wave function.
I switched to the smaller pcS-1 basis set and all that leaking-trouble went away. Everything that we did not expect was gone. TIP3P was converging rather slowly and the PE-model was quickly converging as is shown below for an average of 120 snapshots.
The PE numbers (blue) are converged at QM size 3 but even size 1 and 2 show some good indications. For size 1, acrolein is solvated in purely classical water. As we would expect, the improved potential means that we are converging (fast) towards a value for the absolute shielding of acrolein in water valued -223.2 ppm.
Outlook
We will return to the use of diffuse basis functions and a proper quantum mechanical fix in a later paper as this work here sparked my co-worker +Jógvan Magnus Haugaard Olsen into actually picking up work he had done on repulsion during his Ph.D. (pdf of his thesis)

This work is licensed under a Creative Commons Attribution 3.0 Unported License.
Etiketter:
NMR,
polarizable embedding,
shielding constants
Friday, June 21, 2013
Oxygen, you little rascal!
Calculations of NMR shielding constants using my recent DFT-PE implementation has been underway for some time and I wanted to share the latest results. DFT-PE is a QM/MM type method where the MM region is treated by a polarizable force field calculated for that particular geometry. Initially, I just went away and calculated the NMR shielding constants for acrolein and a few waters with QM and as large an MM region as I could possibly cut out. Unfortunately, the results looked a bit weird when we compared them to a non-polarizable force field (TIP3P) of lesser quality. Lesser in the sense that the description of the electrostatic potential from TIP3P does not match that of the QM electrostatic potential (magnus paper?).
The problem was that the TIP3P results seemed to converge really fast with respect to the size of the QM region and I, using a high-quality PE potential, was converging slower. I went back to the drawing board and came up with the following analysis
Here we see that for increasing sizes of QM region (red) the NMR shielding constant of 17O is reduced by 102 ppm going from acrolein in the gas phase to acrolein surrounded by 53 water molecules (follow the diagonal). Going vertically up in each row, we decrease the size of the QM region, gradually replacing QM waters with MM waters (blue). Deviations from the QM result for that particular system size is shown below each illustration.
Acrolein in 53 MM waters deviates by 23.3 ppm compared to the full QM calculation. This deviation is (if the polarizable force field we are using) purely quantum mechanical. If the first solvation shell is included, that is 7 QM waters, the deviation drops to 4.3 ppm and only improves from here.
Needless to say, TIP3P is also doing great here, but the convergence is not nearly as systematic as is observed for PE. So, it looks like the PE potential is doing it wrong for the right reasons, where as TIP3P is right for the wrong reasons ... that is, using DFT with TIP3P, two wrongs does make a right!
Final note: Oxygen is really, really hard to get right. I showed you only the most difficult case I tried – NMR shielding constants for the rest of acrolein are converged much faster using the smaller QM regions shown above. Luckily for later studies on proteins, nobody does that silly Oxygen atom anyways.

This work is licensed under a Creative Commons Attribution 3.0 Unported License.
Etiketter:
NMR,
pe,
polarizable embedding,
shielding constants
Wednesday, April 10, 2013
NMR shielding constants through polarizable embedding. Part II
In my previous post on the subject of NMR shieldings in polarizable embedding, some progress has been made.
Through the polarizable embedding (PE) approach, calculating gauge-independent magnetic properties in the electrostatic field of the surroundings is now possible. Currently, the PE approach supports an electrostatic multipole expansion up to (and including) order 5, where order 0 is a charge, order 1 is a dipole and so on. The magnetic properties can now be calculated with contributions from quadrupoles (order 2), but we expect to finish the rest of the contributions once some development time is available from the Gen1Int-developers.
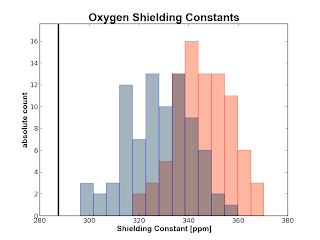
If you have a water cluster and you calculate the shielding constants for the Oxygen and the Hydrogens of the central water molecule with some surrounding water molecules also treated with quantum mechanics, you can now obtain plots like the following
The plots are obtained from 50 snapshots from an MD of water in water. The black vertical line is the experimental number obtained from a (somewhat) recent paper by Kongsted et al. The red histogram is made from numbers calculated at the B3LYP/pcS-0 level of theory whereas the blue histograms are calculated on the same geometry with B3LYP/pcS-1. The gray color in the oxygen plot around the experimental value depicts the uncertainty in that result.
Since this is not going to be a paper on benchmarking your top 20 functionals with your top 5 basis sets, we will from now probably use the KT3 functional with some flavour of the Frank Jensen pcS-n basis sets.
edit 1: Updated the plots to show transparency so you can see both distributions.
edit 2: Added the following based on feedback in the comments to the original post
Janus Eriksen suggested using KT1 or KT2 instead of KT3 since we were only interested in shielding constants. I personally think that Magnus wrote a reasonable response indicating that the difference in using KT1, KT2 and K3 is likely less than 1 ppm in error, but the switch from B3LYP would likely decrease the error by about 13 ppm.
Anders Steen Christen had a valid point in suggesting that we cannot really use the MD-simulations to compare to experiment because the shielding tensors are extremely sensitive to the geometry of the water molecules. Since I haven't really written anything about what we actually aim to do with the data in the end, the point is moot in that respect, but it is a strong point. Perhaps Coupled-Cluster based MD would be the right thing to do.
I would say that this really proves the strength of blogging about your ideas and getting input from others.
Through the polarizable embedding (PE) approach, calculating gauge-independent magnetic properties in the electrostatic field of the surroundings is now possible. Currently, the PE approach supports an electrostatic multipole expansion up to (and including) order 5, where order 0 is a charge, order 1 is a dipole and so on. The magnetic properties can now be calculated with contributions from quadrupoles (order 2), but we expect to finish the rest of the contributions once some development time is available from the Gen1Int-developers.
If you have a water cluster and you calculate the shielding constants for the Oxygen and the Hydrogens of the central water molecule with some surrounding water molecules also treated with quantum mechanics, you can now obtain plots like the following
Since this is not going to be a paper on benchmarking your top 20 functionals with your top 5 basis sets, we will from now probably use the KT3 functional with some flavour of the Frank Jensen pcS-n basis sets.
edit 1: Updated the plots to show transparency so you can see both distributions.
edit 2: Added the following based on feedback in the comments to the original post
Janus Eriksen suggested using KT1 or KT2 instead of KT3 since we were only interested in shielding constants. I personally think that Magnus wrote a reasonable response indicating that the difference in using KT1, KT2 and K3 is likely less than 1 ppm in error, but the switch from B3LYP would likely decrease the error by about 13 ppm.
Anders Steen Christen had a valid point in suggesting that we cannot really use the MD-simulations to compare to experiment because the shielding tensors are extremely sensitive to the geometry of the water molecules. Since I haven't really written anything about what we actually aim to do with the data in the end, the point is moot in that respect, but it is a strong point. Perhaps Coupled-Cluster based MD would be the right thing to do.
I would say that this really proves the strength of blogging about your ideas and getting input from others.
Etiketter:
basis set,
data,
dft,
NMR,
polarizable embedding
Wednesday, March 27, 2013
NMR shielding constants through polarizable embedding
After I finished my Ph.D. in the Jan Jensen group, I've begun working at the University of Southern Denmark with Jacob Kongsted.
Apart from using DALTON instead of GAMESS I've also switched gears and I am now focused on an advanced form of QM/MM which is called polarizable embedding (see this paywalled link for details). Basically your gas-phase Fock operator is extended with interactions from the surroundings. In the language of polarizable embedding (PE) we constructs the effective Fock operator here for Hartree-Fock
$\hat{f}_{eff} = \hat{f}_{HF} + \hat{\nu}_{PE}$
The PE is a sum of interactions from the surroundings onto a molecule of interest. I will deal with the specifics of the PE approach later on.
This is all fine and dandy, but until now the PE model has focused solely on electronic molecular properties. I've extended it so you can calculate nuclear magnetic shielding constants in the PE model using London atomic orbitals (you might know them as Gauge-Including Atomic Orbitals) with contributions from charges, dipoles (induced and static) and quadrupoles. The hope is that we need a small QM region due to a very high-quality potential compared to previous QM/MM studie (see here for an example of needed more than 1000 atoms for converged NMR shieldings).
We'll see how it goes.
Apart from using DALTON instead of GAMESS I've also switched gears and I am now focused on an advanced form of QM/MM which is called polarizable embedding (see this paywalled link for details). Basically your gas-phase Fock operator is extended with interactions from the surroundings. In the language of polarizable embedding (PE) we constructs the effective Fock operator here for Hartree-Fock
$\hat{f}_{eff} = \hat{f}_{HF} + \hat{\nu}_{PE}$
The PE is a sum of interactions from the surroundings onto a molecule of interest. I will deal with the specifics of the PE approach later on.
This is all fine and dandy, but until now the PE model has focused solely on electronic molecular properties. I've extended it so you can calculate nuclear magnetic shielding constants in the PE model using London atomic orbitals (you might know them as Gauge-Including Atomic Orbitals) with contributions from charges, dipoles (induced and static) and quadrupoles. The hope is that we need a small QM region due to a very high-quality potential compared to previous QM/MM studie (see here for an example of needed more than 1000 atoms for converged NMR shieldings).
We'll see how it goes.
Subscribe to:
Posts (Atom)